Difference between revisions of "EpsB"
(→Biological materials) |
|||
| Line 65: | Line 65: | ||
=== Additional information=== | === Additional information=== | ||
| − | |||
| − | |||
| − | |||
=The protein= | =The protein= | ||
| Line 83: | Line 80: | ||
* '''Kinetic information:''' | * '''Kinetic information:''' | ||
| − | * '''Domains:''' | + | * '''[[Domains]]:''' |
* '''Modification:''' | * '''Modification:''' | ||
| − | * ''' | + | * '''[[Cofactors]]:''' |
* '''Effectors of protein activity:''' | * '''Effectors of protein activity:''' | ||
| Line 130: | Line 127: | ||
=Biological materials = | =Biological materials = | ||
| − | * '''Mutant:''' GP1518 (aphA3), available in [[Stülke]] lab | + | * '''Mutant:''' |
| + | ** GP1518 (aphA3), available in [[ Jörg Stülke]]'s lab | ||
| + | ** GP1519 (''[[epsA]]-[[epsB]]'', aphA3), available in [[ Jörg Stülke]]'s lab | ||
* '''Expression vector:''' | * '''Expression vector:''' | ||
| Line 138: | Line 137: | ||
* '''GFP fusion:''' | * '''GFP fusion:''' | ||
| − | * '''two-hybrid system:''' B. pertussis adenylate cyclase-based bacterial two hybrid system ([[BACTH]]), available in [[Stülke]] lab | + | * '''two-hybrid system:''' B. pertussis adenylate cyclase-based bacterial two hybrid system ([[BACTH]]), available in [[ Jörg Stülke]]'s lab |
* '''Antibody:''' | * '''Antibody:''' | ||
| Line 153: | Line 152: | ||
==Reviews== | ==Reviews== | ||
<pubmed>20735481 </pubmed> | <pubmed>20735481 </pubmed> | ||
| − | |||
==Original publications== | ==Original publications== | ||
<pubmed>15661000,16430695,18047568,18647168 18547145 20817675 21856853 21815947 23646920</pubmed> | <pubmed>15661000,16430695,18047568,18647168 18547145 20817675 21856853 21815947 23646920</pubmed> | ||
[[Category:Protein-coding genes]] | [[Category:Protein-coding genes]] | ||
Revision as of 10:27, 31 January 2014
- Description: extracellular polysaccharide synthesis, putative protein tyrosine kinase
| Gene name | epsB |
| Synonyms | yveL |
| Essential | no |
| Product | unknown |
| Function | biofilm formation |
| Gene expression levels in SubtiExpress: epsB | |
| Interactions involving this protein in SubtInteract: EpsB | |
| Regulation of this protein in SubtiPathways: epsB | |
| MW, pI | 24 kDa, 9.918 |
| Gene length, protein length | 681 bp, 227 aa |
| Immediate neighbours | epsC, epsA |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 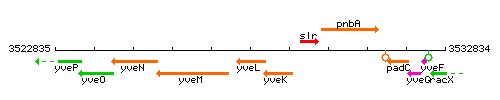
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed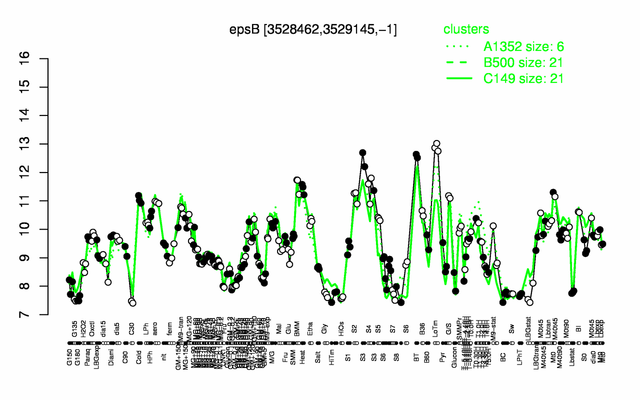
| |
Contents
Categories containing this gene/protein
protein modification, biofilm formation
This gene is a member of the following regulons
AbrB regulon, RemA regulon, SinR regulon
The gene
Basic information
- Locus tag: BSU34360
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: ATP + a [protein]-L-tyrosine = ADP + a [protein]-L-tyrosine phosphate (according to Swiss-Prot)
- Protein family: BY kinase, see the Bacterial Protein Tyrosine Kinase Database)
- Paralogous protein(s): PtkA
Extended information on the protein
- Kinetic information:
- Modification:
- Effectors of protein activity:
Database entries
- UniProt: P71051
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Regulatory mechanism:
- Additional information:
- induction by sequestration of SinR by SinI or SlrA PubMed
- the epsA-epsB-epsC-epsD-epsE-epsF-epsG-epsH-epsI-epsJ-epsK-epsL-epsM-epsN-epsO operon is not expressed in a ymdB mutant PubMed
- the amount of the mRNA is substantially decreased upon depletion of RNase Y (this is likely due to the increased stability of the sinR mRNA) PubMed
Biological materials
- Mutant:
- GP1518 (aphA3), available in Jörg Stülke's lab
- GP1519 (epsA-epsB, aphA3), available in Jörg Stülke's lab
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system: B. pertussis adenylate cyclase-based bacterial two hybrid system (BACTH), available in Jörg Stülke's lab
- Antibody:
- FLAG-tag construct: GP1541 (epsB-3xFLAG spc trpC2) , available in Jörg Stülke's lab
Labs working on this gene/protein
Richard Losick, Harvard Univ., Cambridge, USA homepage
Your additional remarks
References
Reviews
Original publications
Jared T Winkelman, Anna C Bree, Ashley R Bate, Patrick Eichenberger, Richard L Gourse, Daniel B Kearns
RemA is a DNA-binding protein that activates biofilm matrix gene expression in Bacillus subtilis.
Mol Microbiol: 2013, 88(5);984-97
[PubMed:23646920]
[WorldCat.org]
[DOI]
(I p)
Christine Diethmaier, Nico Pietack, Katrin Gunka, Christoph Wrede, Martin Lehnik-Habrink, Christina Herzberg, Sebastian Hübner, Jörg Stülke
A novel factor controlling bistability in Bacillus subtilis: the YmdB protein affects flagellin expression and biofilm formation.
J Bacteriol: 2011, 193(21);5997-6007
[PubMed:21856853]
[WorldCat.org]
[DOI]
(I p)
Martin Lehnik-Habrink, Marc Schaffer, Ulrike Mäder, Christine Diethmaier, Christina Herzberg, Jörg Stülke
RNA processing in Bacillus subtilis: identification of targets of the essential RNase Y.
Mol Microbiol: 2011, 81(6);1459-73
[PubMed:21815947]
[WorldCat.org]
[DOI]
(I p)
Onuma Chumsakul, Hiroki Takahashi, Taku Oshima, Takahiro Hishimoto, Shigehiko Kanaya, Naotake Ogasawara, Shu Ishikawa
Genome-wide binding profiles of the Bacillus subtilis transition state regulator AbrB and its homolog Abh reveals their interactive role in transcriptional regulation.
Nucleic Acids Res: 2011, 39(2);414-28
[PubMed:20817675]
[WorldCat.org]
[DOI]
(I p)
Kazuo Kobayashi
SlrR/SlrA controls the initiation of biofilm formation in Bacillus subtilis.
Mol Microbiol: 2008, 69(6);1399-410
[PubMed:18647168]
[WorldCat.org]
[DOI]
(I p)
Vanesa Olivares-Illana, Philippe Meyer, Emmanuelle Bechet, Virginie Gueguen-Chaignon, Didier Soulat, Sylvie Lazereg-Riquier, Ivan Mijakovic, Josef Deutscher, Alain J Cozzone, Olivier Laprévote, Solange Morera, Christophe Grangeasse, Sylvie Nessler
Structural basis for the regulation mechanism of the tyrosine kinase CapB from Staphylococcus aureus.
PLoS Biol: 2008, 6(6);e143
[PubMed:18547145]
[WorldCat.org]
[DOI]
(I p)
Yunrong Chai, Frances Chu, Roberto Kolter, Richard Losick
Bistability and biofilm formation in Bacillus subtilis.
Mol Microbiol: 2008, 67(2);254-63
[PubMed:18047568]
[WorldCat.org]
[DOI]
(P p)
Frances Chu, Daniel B Kearns, Steven S Branda, Roberto Kolter, Richard Losick
Targets of the master regulator of biofilm formation in Bacillus subtilis.
Mol Microbiol: 2006, 59(4);1216-28
[PubMed:16430695]
[WorldCat.org]
[DOI]
(P p)
Daniel B Kearns, Frances Chu, Steven S Branda, Roberto Kolter, Richard Losick
A master regulator for biofilm formation by Bacillus subtilis.
Mol Microbiol: 2005, 55(3);739-49
[PubMed:15661000]
[WorldCat.org]
[DOI]
(P p)