Difference between revisions of "DivIVA"
(→Biological materials) |
(→Biological materials) |
||
| Line 152: | Line 152: | ||
** GP1482 (chromosomal ''divIVA''-Strep fusion, ''aphA''3), purification from ''B. subtilis'', for [[SPINE]], available in [[Jörg Stülke]]'s lab''' | ** GP1482 (chromosomal ''divIVA''-Strep fusion, ''aphA''3), purification from ''B. subtilis'', for [[SPINE]], available in [[Jörg Stülke]]'s lab''' | ||
| − | * '''Expression vector:''' DivIVA-Strep available [http://www.rki.de/DE/Content/Institut/OrgEinheiten/Abt1/FG11/AG_LM.html here] | + | * '''Expression vector:''' |
| − | + | ** DivIVA-Strep available [http://www.rki.de/DE/Content/Institut/OrgEinheiten/Abt1/FG11/AG_LM.html here] | |
| + | ** pGP1497 (N-terminal Strep-tag fused to C-terminus of ''divIVA'', TEV-site purification from ''E. coli'', in [[pGP172]]), available in [[Jörg Stülke]]'s lab | ||
| + | |||
* '''lacZ fusion:''' | * '''lacZ fusion:''' | ||
Revision as of 13:14, 6 March 2015
- Description: curvature sensitive membrane binding protein that recruits other proteins to the poles and the division septum, cell-division initiation protein (septum placement), part of the Min system (with MinC, MinD, MinJ), Noc and the Min system ensure the efficient utilization of the division site at midcell in by ensuring Z ring placement
| Gene name | divIVA |
| Synonyms | ylmJ |
| Essential | no |
| Product | cell-division initiation protein |
| Function | septum placement |
| Gene expression levels in SubtiExpress: divIVA | |
| Interactions involving this protein in SubtInteract: DivIVA | |
| MW, pI | 19 kDa, 4.846 |
| Gene length, protein length | 492 bp, 164 aa |
| Immediate neighbours | ylmH, ileS |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 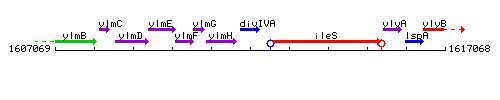
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed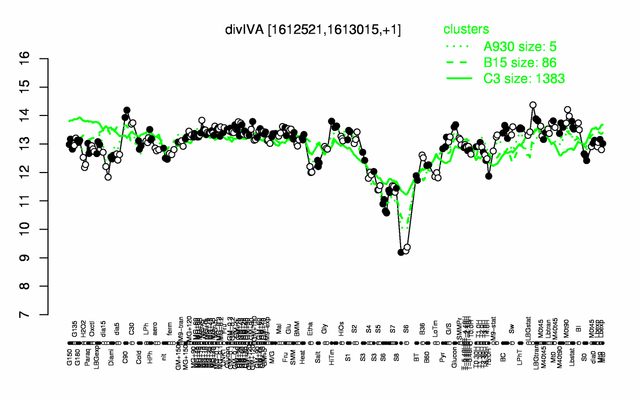
| |
Contents
Categories containing this gene/protein
cell division, membrane proteins, phosphoproteins
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU15420
Phenotypes of a mutant
- Deletion of divIVA leads to filamentation and polar divisions that in turn cause a minicell phenotype. PubMed
- A divIVA mutant has a severe sporulation defect. PubMed
Database entries
- BsubCyc: BSU15420
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
Filamentation is suppressed by mutations in minCD PubMed.
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- curvature sensitive membrane binding protein that recruits other proteins to the poles and the division septum
- DivIVA is required for polar localisation of MinC-MinD via MinJ. PubMed
- It also recruits RacA to the distal pole of the prespore PubMed.
- DivIVA may anchor SpoIIE briefly to the assembling polar septum before SpoIIE is subsequently released into the forespore membrane and recaptured at the polar septum PubMed
- required for the compartment-specific activation of SigF PubMed
- Protein family: gpsB family (according to Swiss-Prot)
- Paralogous protein(s): GpsB
Extended information on the protein
- Kinetic information:
- Modification:
- Cofactors: not known
- Effectors of protein activity: not known
- Localization:
- DivIVA forms a ring underneath the invaginating membrane at the site of cell division and is enriched at both cell poles PubMed.
- forms rings at the division septum and patches at the cell poles PubMed
- membrane targeting requires SecA PubMed
- assembles into a ring-like structure at the polar septum during sporulation PubMed
Database entries
- BsubCyc: BSU15420
- UniProt: P71021
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Operon: divIVA PubMed
- Regulation:
- Additional information:
Biological materials
- Mutant:
- 4041 (divIVA::tet), available in Leendert Hamoen's, Jörg Stülke's, and Sven Halbedel 's lab
- GP1482 (chromosomal divIVA-Strep fusion, aphA3), purification from B. subtilis, for SPINE, available in Jörg Stülke's lab
- Expression vector:
- DivIVA-Strep available here
- pGP1497 (N-terminal Strep-tag fused to C-terminus of divIVA, TEV-site purification from E. coli, in pGP172), available in Jörg Stülke's lab
- lacZ fusion:
- GFP fusion: divIVA-gfp fusions available from the Hamoen Lab
- two-hybrid system: B. pertussis adenylate cyclase-based bacterial two hybrid system (BACTH), available in Sven Halbedel's and Jörg Stülke's labs
- Antibody: A polyclonal anti-DivIVA antiserum generated in rabbit is described here PubMed.
Labs working on this gene/protein
Leendert Hamoen, Centre for Bacterial Cell Biology, Newcastle upon Tyne, United Kingdom [4]
Imrich Barak, Slovak Academy of Science, Bratislava, Slovakia homepage
Sven Halbedel, Robert Koch Institute homepage
Your additional remarks
References
Reviews
Original Publications
Prahathees Eswaramoorthy, Peter W Winter, Peter Wawrzusin, Andrew G York, Hari Shroff, Kumaran S Ramamurthi
Asymmetric division and differential gene expression during a bacterial developmental program requires DivIVA.
PLoS Genet: 2014, 10(8);e1004526
[PubMed:25101664]
[WorldCat.org]
[DOI]
(I e)
Juri N Bach, Nadine Albrecht, Marc Bramkamp
Imaging DivIVA dynamics using photo-convertible and activatable fluorophores in Bacillus subtilis.
Front Microbiol: 2014, 5;59
[PubMed:24600441]
[WorldCat.org]
[DOI]
(P e)
Sven Halbedel, Maki Kawai, Reinhard Breitling, Leendert W Hamoen
SecA is required for membrane targeting of the cell division protein DivIVA in vivo.
Front Microbiol: 2014, 5;58
[PubMed:24592260]
[WorldCat.org]
[DOI]
(P e)
Lars D Renner, Prahathees Eswaramoorthy, Kumaran S Ramamurthi, Douglas B Weibel
Studying biomolecule localization by engineering bacterial cell wall curvature.
PLoS One: 2013, 8(12);e84143
[PubMed:24391905]
[WorldCat.org]
[DOI]
(I e)
Monika Vishnoi, Jatin Narula, Seram Nganbiton Devi, Hoang-Anh Dao, Oleg A Igoshin, Masaya Fujita
Triggering sporulation in Bacillus subtilis with artificial two-component systems reveals the importance of proper Spo0A activation dynamics.
Mol Microbiol: 2013, 90(1);181-94
[PubMed:23927765]
[WorldCat.org]
[DOI]
(I p)
Erik Nico Trip, Jan-Willem Veening, Eric J Stewart, Jeff Errington, Dirk-Jan Scheffers
Balanced transcription of cell division genes in Bacillus subtilis as revealed by single cell analysis.
Environ Microbiol: 2013, 15(12);3196-209
[PubMed:23701187]
[WorldCat.org]
[DOI]
(I p)
Suey van Baarle, Ilkay Nazli Celik, Karan Gautam Kaval, Marc Bramkamp, Leendert W Hamoen, Sven Halbedel
Protein-protein interaction domains of Bacillus subtilis DivIVA.
J Bacteriol: 2013, 195(5);1012-21
[PubMed:23264578]
[WorldCat.org]
[DOI]
(I p)
Joe Pogliano, Nicolas Pogliano, Jared A Silverman
Daptomycin-mediated reorganization of membrane architecture causes mislocalization of essential cell division proteins.
J Bacteriol: 2012, 194(17);4494-504
[PubMed:22661688]
[WorldCat.org]
[DOI]
(I p)
Valquiria Tiago dos Santos, Alexandre W Bisson-Filho, Frederico J Gueiros-Filho
DivIVA-mediated polar localization of ComN, a posttranscriptional regulator of Bacillus subtilis.
J Bacteriol: 2012, 194(14);3661-9
[PubMed:22582279]
[WorldCat.org]
[DOI]
(I p)
Alexander K W Elsholz, Kürsad Turgay, Stephan Michalik, Bernd Hessling, Katrin Gronau, Dan Oertel, Ulrike Mäder, Jörg Bernhardt, Dörte Becher, Michael Hecker, Ulf Gerth
Global impact of protein arginine phosphorylation on the physiology of Bacillus subtilis.
Proc Natl Acad Sci U S A: 2012, 109(19);7451-6
[PubMed:22517742]
[WorldCat.org]
[DOI]
(I p)
Christopher D A Rodrigues, Elizabeth J Harry
The Min system and nucleoid occlusion are not required for identifying the division site in Bacillus subtilis but ensure its efficient utilization.
PLoS Genet: 2012, 8(3);e1002561
[PubMed:22457634]
[WorldCat.org]
[DOI]
(I p)
Prahathees Eswaramoorthy, Marcella L Erb, James A Gregory, Jared Silverman, Kit Pogliano, Joe Pogliano, Kumaran S Ramamurthi
Cellular architecture mediates DivIVA ultrastructure and regulates min activity in Bacillus subtilis.
mBio: 2011, 2(6);
[PubMed:22108385]
[WorldCat.org]
[DOI]
(I e)
Kenneth Briley, Peter Prepiak, Miguel J Dias, Jeanette Hahn, David Dubnau
Maf acts downstream of ComGA to arrest cell division in competent cells of B. subtilis.
Mol Microbiol: 2011, 81(1);23-39
[PubMed:21564336]
[WorldCat.org]
[DOI]
(I p)
Maria A Oliva, Sven Halbedel, Stefan M Freund, Pavel Dutow, Thomas A Leonard, Dmitry B Veprintsev, Leendert W Hamoen, Jan Löwe
Features critical for membrane binding revealed by DivIVA crystal structure.
EMBO J: 2010, 29(12);1988-2001
[PubMed:20502438]
[WorldCat.org]
[DOI]
(I p)
Suey van Baarle, Marc Bramkamp
The MinCDJ system in Bacillus subtilis prevents minicell formation by promoting divisome disassembly.
PLoS One: 2010, 5(3);e9850
[PubMed:20352045]
[WorldCat.org]
[DOI]
(I e)
Kumaran S Ramamurthi, Richard Losick
Negative membrane curvature as a cue for subcellular localization of a bacterial protein.
Proc Natl Acad Sci U S A: 2009, 106(32);13541-5
[PubMed:19666580]
[WorldCat.org]
[DOI]
(I p)
Jennifer R Juarez, William Margolin
Irresistible curves.
EMBO J: 2009, 28(15);2147-8
[PubMed:19654604]
[WorldCat.org]
[DOI]
(I p)
Rok Lenarcic, Sven Halbedel, Loek Visser, Michael Shaw, Ling Juan Wu, Jeff Errington, Davide Marenduzzo, Leendert W Hamoen
Localisation of DivIVA by targeting to negatively curved membranes.
EMBO J: 2009, 28(15);2272-82
[PubMed:19478798]
[WorldCat.org]
[DOI]
(I p)
Pamela Gamba, Jan-Willem Veening, Nigel J Saunders, Leendert W Hamoen, Richard A Daniel
Two-step assembly dynamics of the Bacillus subtilis divisome.
J Bacteriol: 2009, 191(13);4186-94
[PubMed:19429628]
[WorldCat.org]
[DOI]
(I p)
Marc Bramkamp, Robyn Emmins, Louise Weston, Catriona Donovan, Richard A Daniel, Jeff Errington
A novel component of the division-site selection system of Bacillus subtilis and a new mode of action for the division inhibitor MinCD.
Mol Microbiol: 2008, 70(6);1556-69
[PubMed:19019154]
[WorldCat.org]
[DOI]
(I p)
S E Perry, D H Edwards
Identification of a polar targeting determinant for Bacillus subtilis DivIVA.
Mol Microbiol: 2004, 54(5);1237-49
[PubMed:15554965]
[WorldCat.org]
[DOI]
(P p)
Virginie Molle, Masaya Fujita, Shane T Jensen, Patrick Eichenberger, José E González-Pastor, Jun S Liu, Richard Losick
The Spo0A regulon of Bacillus subtilis.
Mol Microbiol: 2003, 50(5);1683-701
[PubMed:14651647]
[WorldCat.org]
[DOI]
(P p)
Leendert W Hamoen, Jeffery Errington
Polar targeting of DivIVA in Bacillus subtilis is not directly dependent on FtsZ or PBP 2B.
J Bacteriol: 2003, 185(2);693-7
[PubMed:12511520]
[WorldCat.org]
[DOI]
(P p)
Frederico J Gueiros-Filho, Richard Losick
A widely conserved bacterial cell division protein that promotes assembly of the tubulin-like protein FtsZ.
Genes Dev: 2002, 16(19);2544-56
[PubMed:12368265]
[WorldCat.org]
[DOI]
(P p)
M E Karoui, J Errington
Isolation and characterization of topological specificity mutants of minD in Bacillus subtilis.
Mol Microbiol: 2001, 42(5);1211-21
[PubMed:11886553]
[WorldCat.org]
[DOI]
(P p)
H B Thomaides, M Freeman, M El Karoui, J Errington
Division site selection protein DivIVA of Bacillus subtilis has a second distinct function in chromosome segregation during sporulation.
Genes Dev: 2001, 15(13);1662-73
[PubMed:11445541]
[WorldCat.org]
[DOI]
(P p)
D H Edwards, H B Thomaides, J Errington
Promiscuous targeting of Bacillus subtilis cell division protein DivIVA to division sites in Escherichia coli and fission yeast.
EMBO J: 2000, 19(11);2719-27
[PubMed:10835369]
[WorldCat.org]
[DOI]
(P p)
D H Edwards, J Errington
The Bacillus subtilis DivIVA protein targets to the division septum and controls the site specificity of cell division.
Mol Microbiol: 1997, 24(5);905-15
[PubMed:9219999]
[WorldCat.org]
[DOI]
(P p)
J H Cha, G C Stewart
The divIVA minicell locus of Bacillus subtilis.
J Bacteriol: 1997, 179(5);1671-83
[PubMed:9045828]
[WorldCat.org]
[DOI]
(P p)