Difference between revisions of "NtdB"
| Line 1: | Line 1: | ||
| − | + | ||
{| align="right" border="1" cellpadding="2" | {| align="right" border="1" cellpadding="2" | ||
| Line 13: | Line 13: | ||
|- | |- | ||
|style="background:#ABCDEF;" align="center"|'''Function''' || synthesis of the antibiotic neotrehalosadiamine | |style="background:#ABCDEF;" align="center"|'''Function''' || synthesis of the antibiotic neotrehalosadiamine | ||
| + | |- | ||
| + | |colspan="2" style="background:#FAF8CC;" align="center"| '''Gene expression levels in [http://cellpublisher.gobics.de/subtiexpress/ ''Subti''Express]''': [http://cellpublisher.gobics.de/subtiexpress/bsu/BSU10540 ntdB] | ||
|- | |- | ||
|style="background:#ABCDEF;" align="center"| '''MW, pI''' || 33 kDa, 5.352 | |style="background:#ABCDEF;" align="center"| '''MW, pI''' || 33 kDa, 5.352 | ||
Revision as of 16:32, 6 August 2012
| Gene name | ntdB |
| Synonyms | yhjK |
| Essential | no |
| Product | unknown |
| Function | synthesis of the antibiotic neotrehalosadiamine |
| Gene expression levels in SubtiExpress: ntdB | |
| MW, pI | 33 kDa, 5.352 |
| Gene length, protein length | 858 bp, 286 aa |
| Immediate neighbours | ntdC, ntdA |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 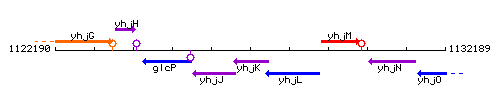
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed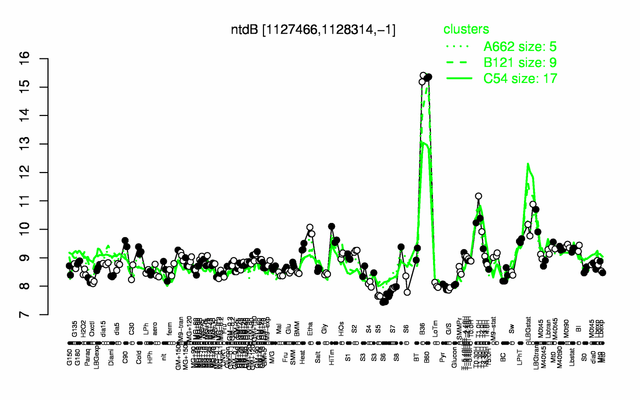
| |
Contents
Categories containing this gene/protein
miscellaneous metabolic pathways, biosynthesis of antibacterial compounds
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU10540
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family: Cof family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
Database entries
- Structure: 3GYG
- UniProt: O07565
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Reviews
Original publications
Takashi Inaoka, Takenori Satomura, Yasutaro Fujita, Kozo Ochi
Novel gene regulation mediated by overproduction of secondary metabolite neotrehalosadiamine in Bacillus subtilis.
FEMS Microbiol Lett: 2009, 291(2);151-6
[PubMed:19087206]
[WorldCat.org]
[DOI]
(I p)
Takashi Inaoka, Kozo Ochi
Glucose uptake pathway-specific regulation of synthesis of neotrehalosadiamine, a novel autoinducer produced in Bacillus subtilis.
J Bacteriol: 2007, 189(1);65-75
[PubMed:17056753]
[WorldCat.org]
[DOI]
(P p)
Takashi Inaoka, Kosaku Takahashi, Hiroshi Yada, Mitsuru Yoshida, Kozo Ochi
RNA polymerase mutation activates the production of a dormant antibiotic 3,3'-neotrehalosadiamine via an autoinduction mechanism in Bacillus subtilis.
J Biol Chem: 2004, 279(5);3885-92
[PubMed:14612444]
[WorldCat.org]
[DOI]
(P p)