Difference between revisions of "CitB"
| Line 31: | Line 31: | ||
<div align="right"> <small>This image was kindly provided by [http://genolist.pasteur.fr/SubtiList/ SubtiList]</small></div> | <div align="right"> <small>This image was kindly provided by [http://genolist.pasteur.fr/SubtiList/ SubtiList]</small></div> | ||
|- | |- | ||
| − | |colspan="2" |'''[http://genome.jouy.inra.fr/cgi-bin/seb/viewdetail.py?id=citB_1926680_1929409_1 Expression at a glance]'''   {{PubMed|22383849}}<br/>[[Image:citB_expression.png|500px]] | + | |colspan="2" |'''[http://genome.jouy.inra.fr/cgi-bin/seb/viewdetail.py?id=citB_1926680_1929409_1 Expression at a glance]'''   {{PubMed|22383849}}<br/>[[Image:citB_expression.png|500px|link=http://subtiwiki.uni-goettingen.de/apps/expression/expression.php?search=BSU18000]] |
|- | |- | ||
|} | |} | ||
Revision as of 13:29, 16 May 2013
- Description: trigger enzyme: aconitase and RNA binding protein
| Gene name | citB |
| Synonyms | |
| Essential | no |
| Product | trigger enzyme: aconitate hydratase (aconitase) |
| Function | TCA cycle |
| Gene expression levels in SubtiExpress: citB | |
| Interactions involving this protein in SubtInteract: CitB | |
| Metabolic function and regulation of this protein in SubtiPathways: Central C-metabolism | |
| MW, pI | 99 kDa, 4.903 |
| Gene length, protein length | 2727 bp, 909 aa |
| Immediate neighbours | sspO, yneN |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 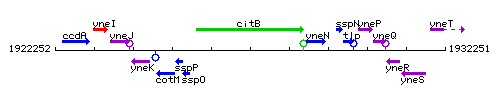
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed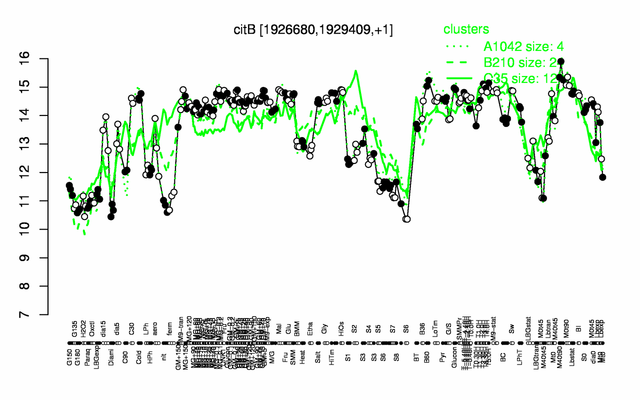
| |
Contents
Categories containing this gene/protein
carbon core metabolism, trigger enzyme, RNA binding regulators
This gene is a member of the following regulons
CcpA regulon, CcpC regulon, CodY regulon, FsrA regulon
The CitB regulon: feuA-feuB-feuC-ybbA, citZ
The gene
Basic information
- Locus tag: BSU18000
Phenotypes of a mutant
- glutamate auxotrophy and a defect in sporulation PubMed
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
- A mutation was found in this gene after evolution under relaxed selection for sporulation PubMed
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Citrate <=> isocitrate
- Binding to iron responsive elements (IRE RNA) in the absence of the FeS cluster PubMed
- Protein family:
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s): FeS cluster
- Effectors of protein activity:
Database entries
- Structure: 1L5J (E. coli)
- UniProt: P09339
- KEGG entry: [3]
- E.C. number: 4.2.1.3
Additional information
- B. subtilis aconitase is both an enzyme and an RNA binding protein (moonlighting protein) PubMed
- extensive information on the structure and enzymatic properties of CitB can be found at Proteopedia
Expression and regulation
- Regulation:
- repressed during growth in the presence of branched chain amino acids (CodY) PubMed
- repressed in the presence of glucose and glutamate (CcpC) PubMed
- expressed upon transition into the stationary phase (AbrB) PubMed, indirect negative regulation by AbrB PubMed
- repressed by glucose (3.7-fold) (CcpA) PubMed
- repression by glucose + arginine (CcpC) PubMed
- less expressed under conditions of extreme iron limitation (FsrA) PubMed
- part of the iron sparing response (FsrA) PubMed
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant:
- GP683 (erm), available in Jörg Stülke's lab
- GP1441 (spc), available in Jörg Stülke's lab
- 1A999 ( citB::spec), PubMed, available at BGSC
- Expression vector:
- GP1439 (citB-Strep (spc)), purification from B. subtilis, for SPINE, available in Jörg Stülke's lab
- pGP1810 (for expression, purification in E. coli with N-terminal Strep-tag, in pGP172, available in Jörg Stülke's lab
- lacZ fusion:
- pGP700 (in pAC5), available in Jörg Stülke's lab
- GFP fusion: GP1434 (spc, based on pGP1870), available in Jörg Stülke's lab
- two-hybrid system: B. pertussis adenylate cyclase-based bacterial two hybrid system (BACTH), available in Jörg Stülke's lab
- Antibody: available in Linc Sonenshein's lab
- FLAG-tag construct:
- GP1144 (spc, based on pGP1331), available in Jörg Stülke's lab
- GP1145 (kan), available in Jörg Stülke's lab
Labs working on this gene/protein
- Linc Sonenshein, Tufts University, Boston, MA, USA Homepage
- Jörg Stülke, University of Göttingen, Germany Homepage
Your additional remarks
References
Reviews
Original publications
Additional publications: PubMed
Kieran B Pechter, Frederik M Meyer, Alisa W Serio, Jörg Stülke, Abraham L Sonenshein
Two roles for aconitase in the regulation of tricarboxylic acid branch gene expression in Bacillus subtilis.
J Bacteriol: 2013, 195(7);1525-37
[PubMed:23354745]
[WorldCat.org]
[DOI]
(I p)
Meghna Mittal, Kieran B Pechter, Silvia Picossi, Hyun-Jin Kim, Kathryn O Kerstein, Abraham L Sonenshein
Dual role of CcpC protein in regulation of aconitase gene expression in Listeria monocytogenes and Bacillus subtilis.
Microbiology (Reading): 2013, 159(Pt 1);68-76
[PubMed:23139400]
[WorldCat.org]
[DOI]
(I p)
Gregory T Smaldone, Olga Revelles, Ahmed Gaballa, Uwe Sauer, Haike Antelmann, John D Helmann
A global investigation of the Bacillus subtilis iron-sparing response identifies major changes in metabolism.
J Bacteriol: 2012, 194(10);2594-605
[PubMed:22389480]
[WorldCat.org]
[DOI]
(I p)
Christopher T Brown, Laura K Fishwick, Binna M Chokshi, Marissa A Cuff, Jay M Jackson, Travis Oglesby, Alison T Rioux, Enrique Rodriguez, Gregory S Stupp, Austin H Trupp, James S Woollcombe-Clarke, Tracy N Wright, William J Zaragoza, Jennifer C Drew, Eric W Triplett, Wayne L Nicholson
Whole-genome sequencing and phenotypic analysis of Bacillus subtilis mutants following evolution under conditions of relaxed selection for sporulation.
Appl Environ Microbiol: 2011, 77(19);6867-77
[PubMed:21821766]
[WorldCat.org]
[DOI]
(I p)
Weihua Gao, Sen Dai, Quanli Liu, Haijin Xu, Yanlin Bai, Mingqiang Qiao
Effect of site-directed mutagenesis of citB on the expression and activity of Bacillus subtilis aconitase.
Mikrobiologiia: 2010, 79(6);774-8
[PubMed:21446632]
[WorldCat.org]
(P p)
Weihua Gao, Sen Dai, Quanli Liu, Haijin Xu, Mingqiang Qiao
CitB mutation increases the alkaline protease productivity in Bacillus subtilis.
J Gen Appl Microbiol: 2010, 56(5);403-7
[PubMed:21099137]
[WorldCat.org]
[DOI]
(P p)
Frederik M Meyer, Jan Gerwig, Elke Hammer, Christina Herzberg, Fabian M Commichau, Uwe Völker, Jörg Stülke
Physical interactions between tricarboxylic acid cycle enzymes in Bacillus subtilis: evidence for a metabolon.
Metab Eng: 2011, 13(1);18-27
[PubMed:20933603]
[WorldCat.org]
[DOI]
(I p)
Alexander G Albrecht, Daili J A Netz, Marcus Miethke, Antonio J Pierik, Olaf Burghaus, Florian Peuckert, Roland Lill, Mohamed A Marahiel
SufU is an essential iron-sulfur cluster scaffold protein in Bacillus subtilis.
J Bacteriol: 2010, 192(6);1643-51
[PubMed:20097860]
[WorldCat.org]
[DOI]
(I p)
Ahmed Gaballa, Haike Antelmann, Claudio Aguilar, Sukhjit K Khakh, Kyung-Bok Song, Gregory T Smaldone, John D Helmann
The Bacillus subtilis iron-sparing response is mediated by a Fur-regulated small RNA and three small, basic proteins.
Proc Natl Acad Sci U S A: 2008, 105(33);11927-32
[PubMed:18697947]
[WorldCat.org]
[DOI]
(I p)
Alisa W Serio, Kieran B Pechter, Abraham L Sonenshein
Bacillus subtilis aconitase is required for efficient late-sporulation gene expression.
J Bacteriol: 2006, 188(17);6396-405
[PubMed:16923907]
[WorldCat.org]
[DOI]
(P p)
Hans-Matti Blencke, Irene Reif, Fabian M Commichau, Christian Detsch, Ingrid Wacker, Holger Ludwig, Jörg Stülke
Regulation of citB expression in Bacillus subtilis: integration of multiple metabolic signals in the citrate pool and by the general nitrogen regulatory system.
Arch Microbiol: 2006, 185(2);136-46
[PubMed:16395550]
[WorldCat.org]
[DOI]
(P p)
Hans-Matti Blencke, Georg Homuth, Holger Ludwig, Ulrike Mäder, Michael Hecker, Jörg Stülke
Transcriptional profiling of gene expression in response to glucose in Bacillus subtilis: regulation of the central metabolic pathways.
Metab Eng: 2003, 5(2);133-49
[PubMed:12850135]
[WorldCat.org]
[DOI]
(P p)
Hyun-Jin Kim, Sam-In Kim, Manoja Ratnayake-Lecamwasam, Kiyoshi Tachikawa, Abraham L Sonenshein, Mark Strauch
Complex regulation of the Bacillus subtilis aconitase gene.
J Bacteriol: 2003, 185(5);1672-80
[PubMed:12591885]
[WorldCat.org]
[DOI]
(P p)
C Jourlin-Castelli, N Mani, M M Nakano, A L Sonenshein
CcpC, a novel regulator of the LysR family required for glucose repression of the citB gene in Bacillus subtilis.
J Mol Biol: 2000, 295(4);865-78
[PubMed:10656796]
[WorldCat.org]
[DOI]
(P p)
C Alén, A L Sonenshein
Bacillus subtilis aconitase is an RNA-binding protein.
Proc Natl Acad Sci U S A: 1999, 96(18);10412-7
[PubMed:10468622]
[WorldCat.org]
[DOI]
(P p)
M M Nakano, P Zuber, A L Sonenshein
Anaerobic regulation of Bacillus subtilis Krebs cycle genes.
J Bacteriol: 1998, 180(13);3304-11
[PubMed:9642180]
[WorldCat.org]
[DOI]
(P p)
J E Craig, M J Ford, D C Blaydon, A L Sonenshein
A null mutation in the Bacillus subtilis aconitase gene causes a block in Spo0A-phosphate-dependent gene expression.
J Bacteriol: 1997, 179(23);7351-9
[PubMed:9393699]
[WorldCat.org]
[DOI]
(P p)
A Fouet, S F Jin, G Raffel, A L Sonenshein
Multiple regulatory sites in the Bacillus subtilis citB promoter region.
J Bacteriol: 1990, 172(9);5408-15
[PubMed:2118511]
[WorldCat.org]
[DOI]
(P p)
A Fouet, A L Sonenshein
A target for carbon source-dependent negative regulation of the citB promoter of Bacillus subtilis.
J Bacteriol: 1990, 172(2);835-44
[PubMed:2105305]
[WorldCat.org]
[DOI]
(P p)
M S Rosenkrantz, D W Dingman, A L Sonenshein
Bacillus subtilis citB gene is regulated synergistically by glucose and glutamine.
J Bacteriol: 1985, 164(1);155-64
[PubMed:2413006]
[WorldCat.org]
[DOI]
(P p)
S H Fisher, B Magasanik
2-Ketoglutarate and the regulation of aconitase and histidase formation in Bacillus subtilis.
J Bacteriol: 1984, 158(1);379-82
[PubMed:6143742]
[WorldCat.org]
[DOI]
(P p)