Difference between revisions of "SigD"
(→Expression and regulation) |
|||
| Line 111: | Line 111: | ||
=Expression and regulation= | =Expression and regulation= | ||
| − | * '''Operon:''' | + | * '''Operon:''' part of the [[fla-che operon]] |
** ''[[flgB]]-[[flgC]]-[[fliE]]-[[fliF]]-[[fliG]]-[[fliH]]-[[fliI]]-[[fliJ]]-[[ylxF]]-[[fliK]]-[[flgD]]-[[flgE]]-[[fliL]]-[[fliM]]-[[fliY]]-[[cheY]]-[[fliZ]]-[[fliP]]-[[fliQ]]-[[fliR]]-[[flhB]]-[[flhA]]-[[flhF]]-[[flhG]]-[[cheB]]-[[cheA]]-[[cheW]]-[[cheC]]-[[cheD]]-[[sigD]]-[[swrB]]'' {{PubMed|9657996,8157612,15175317}} | ** ''[[flgB]]-[[flgC]]-[[fliE]]-[[fliF]]-[[fliG]]-[[fliH]]-[[fliI]]-[[fliJ]]-[[ylxF]]-[[fliK]]-[[flgD]]-[[flgE]]-[[fliL]]-[[fliM]]-[[fliY]]-[[cheY]]-[[fliZ]]-[[fliP]]-[[fliQ]]-[[fliR]]-[[flhB]]-[[flhA]]-[[flhF]]-[[flhG]]-[[cheB]]-[[cheA]]-[[cheW]]-[[cheC]]-[[cheD]]-[[sigD]]-[[swrB]]'' {{PubMed|9657996,8157612,15175317}} | ||
** ''[[ylxF]]-[[fliK]]-[[flgD]]-[[flgE]]-[[fliL]]-[[fliM]]-[[fliY]]-[[cheY]]-[[fliZ]]-[[fliP]]-[[fliQ]]-[[fliR]]-[[flhB]]-[[flhA]]-[[flhF]]-[[flhG]]-[[cheB]]-[[cheA]]-[[cheW]]-[[cheC]]-[[cheD]]-[[sigD]]-[[swrB]]'' {{PubMed|20233303}} | ** ''[[ylxF]]-[[fliK]]-[[flgD]]-[[flgE]]-[[fliL]]-[[fliM]]-[[fliY]]-[[cheY]]-[[fliZ]]-[[fliP]]-[[fliQ]]-[[fliR]]-[[flhB]]-[[flhA]]-[[flhF]]-[[flhG]]-[[cheB]]-[[cheA]]-[[cheW]]-[[cheC]]-[[cheD]]-[[sigD]]-[[swrB]]'' {{PubMed|20233303}} | ||
| Line 123: | Line 123: | ||
** ''[[sigD]]'': [[SigA]] {{PubMed|9335309}} | ** ''[[sigD]]'': [[SigA]] {{PubMed|9335309}} | ||
| − | * '''Regulation:''' | + | * '''Regulation:''' see [[fla-che operon]] |
| − | |||
| − | * '''Regulatory mechanism:''' | + | * '''Regulatory mechanism:''' see [[fla-che operon]] |
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
* '''Additional information:''' | * '''Additional information:''' | ||
Revision as of 14:18, 12 February 2015
- Description: RNA polymerase sigma factor SigD
| Gene name | sigD |
| Synonyms | flaB |
| Essential | no |
| Product | RNA polymerase sigma factor SigD |
| Function | regulation of flagella, motility, chemotaxis and autolysis |
| Gene expression levels in SubtiExpress: sigD | |
| Interactions involving this protein in SubtInteract: SigD | |
| MW, pI | 29 kDa, 5.167 |
| Gene length, protein length | 762 bp, 254 aa |
| Immediate neighbours | cheD, swrB |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 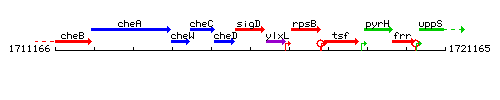
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed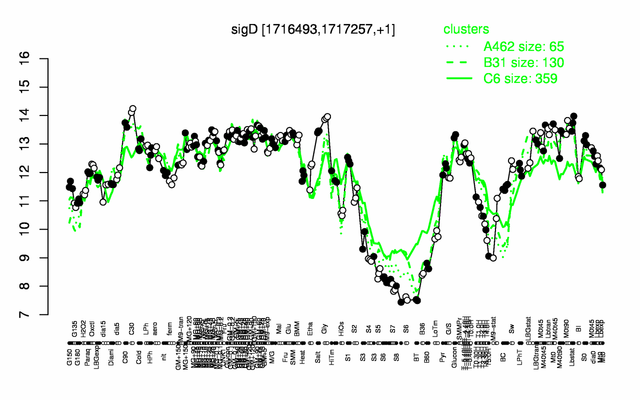
| |
Contents
Categories containing this gene/protein
transcription, sigma factors and their control, motility and chemotaxis
This gene is a member of the following regulons
CodY regulon, DegU regulon, SigD regulon, Spo0A regulon
The SigD regulon
The gene
Basic information
- Locus tag: BSU16470
Phenotypes of a mutant
- Inactivation of alsR, sigD and sigW increases competitive fitness of Bacillus subtilis under non-sporulating conditions PubMed
- mucoid phenotype due to the overproduction of poly-gamma-glutamate (because of the loss of motA-motB expression) PubMed
Database entries
- BsubCyc: BSU16470
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family: sigma-70 factor family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
Database entries
- BsubCyc: BSU16470
- Structure:
- UniProt: P10726
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Operon: part of the fla-che operon
- flgB-flgC-fliE-fliF-fliG-fliH-fliI-fliJ-ylxF-fliK-flgD-flgE-fliL-fliM-fliY-cheY-fliZ-fliP-fliQ-fliR-flhB-flhA-flhF-flhG-cheB-cheA-cheW-cheC-cheD-sigD-swrB PubMed
- ylxF-fliK-flgD-flgE-fliL-fliM-fliY-cheY-fliZ-fliP-fliQ-fliR-flhB-flhA-flhF-flhG-cheB-cheA-cheW-cheC-cheD-sigD-swrB PubMed
- sigD-swrB PubMed
- Regulation: see fla-che operon
- Regulatory mechanism: see fla-che operon
- Additional information:
Biological materials
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system: B. pertussis adenylate cyclase-based bacterial two hybrid system (BACTH), available in Stülke lab
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Reviews
Sampriti Mukherjee, Daniel B Kearns
The structure and regulation of flagella in Bacillus subtilis.
Annu Rev Genet: 2014, 48;319-40
[PubMed:25251856]
[WorldCat.org]
[DOI]
(I p)
Jan-Willem Veening, Oscar P Kuipers
Gene position within a long transcript as a determinant for stochastic switching in bacteria.
Mol Microbiol: 2010, 76(2);269-72
[PubMed:20233302]
[WorldCat.org]
[DOI]
(I p)
The SigD regulon
Original publications