Difference between revisions of "GlnA"
| Line 1: | Line 1: | ||
| − | + | * '''Description:''' [[trigger enzymes|trigger enzyme]]: glutamine synthetase and effector of [[TnrA]] and [[GlnR]] <br/><br/> | |
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | * '''Description:''' [[trigger enzymes|trigger enzyme]]: glutamine synthetase and effector of [[TnrA]] and [[GlnR]] | ||
{| align="right" border="1" cellpadding="2" | {| align="right" border="1" cellpadding="2" | ||
| Line 22: | Line 14: | ||
|style="background:#ABCDEF;" align="center"|'''Function''' || glutamine biosynthesis, control of [[TnrA]] and [[GlnR]] activity | |style="background:#ABCDEF;" align="center"|'''Function''' || glutamine biosynthesis, control of [[TnrA]] and [[GlnR]] activity | ||
|- | |- | ||
| − | |colspan="2" style="background:#FAF8CC;" align="center"| '''Metabolic function and regulation of this protein in [[SubtiPathways|''Subti''Pathways]]: | + | |colspan="2" style="background:#FAF8CC;" align="center"| '''Metabolic function and regulation of this protein in [[SubtiPathways|''Subti''Pathways]]: <br/>[http://subtiwiki.uni-goettingen.de/pathways/glutamate.html Ammonium/ glutamate]''' |
|- | |- | ||
|style="background:#ABCDEF;" align="center"| '''MW, pI''' || 50 kDa, 4.874 | |style="background:#ABCDEF;" align="center"| '''MW, pI''' || 50 kDa, 4.874 | ||
| Line 30: | Line 22: | ||
|style="background:#ABCDEF;" align="center"|'''Immediate neighbours''' || ''[[glnR]]'', ''[[ynxB]]'' | |style="background:#ABCDEF;" align="center"|'''Immediate neighbours''' || ''[[glnR]]'', ''[[ynxB]]'' | ||
|- | |- | ||
| − | |colspan="2" style="background:#FAF8CC;" align="center"|'''Get the DNA and protein [http://srs.ebi.ac.uk/srsbin/cgi-bin/wgetz?-e+& | + | |colspan="2" style="background:#FAF8CC;" align="center"|'''Get the DNA and protein [http://srs.ebi.ac.uk/srsbin/cgi-bin/wgetz?-e+[EMBLCDS:CAB13630]+-newId sequences] <br/> (Barbe ''et al.'', 2009)''' |
|- | |- | ||
| − | |colspan="2" | '''Genetic context''' | + | |colspan="2" | '''Genetic context''' <br/> [[Image:glnA_context.gif]] |
| − | + | <div align="right"> <small>This image was kindly provided by [http://genolist.pasteur.fr/SubtiList/ SubtiList]</small></div> | |
|- | |- | ||
|} | |} | ||
| Line 39: | Line 31: | ||
__TOC__ | __TOC__ | ||
| − | + | <br/><br/><br/><br/><br/><br/> | |
=The gene= | =The gene= | ||
| Line 64: | Line 56: | ||
=== Basic information/ Evolution === | === Basic information/ Evolution === | ||
| − | * '''Catalyzed reaction/ biological activity:''' ATP + L-glutamate + NH | + | * '''Catalyzed reaction/ biological activity:''' ATP + L-glutamate + NH<sub>3</sub> = ADP + phosphate + L-glutamine (according to Swiss-Prot) |
* '''Protein family:''' glutamine synthetase family (according to Swiss-Prot) | * '''Protein family:''' glutamine synthetase family (according to Swiss-Prot) | ||
| Line 139: | Line 131: | ||
=References= | =References= | ||
==Reviews== | ==Reviews== | ||
| − | + | <pubmed> 10231480 18086213 </pubmed> | |
==Original publications== | ==Original publications== | ||
| − | + | <pubmed>19233925, 20389117,8799114,18195355, 11719184, 12139611, 2573733, 8636055, 19233925, 16493705, 16885465, 6141156 2906311 20656908 16055443 18331450 16547045 8093698 </pubmed> | |
[[Category:Protein-coding genes]] | [[Category:Protein-coding genes]] | ||
Revision as of 04:58, 24 November 2010
- Description: trigger enzyme: glutamine synthetase and effector of TnrA and GlnR
| Gene name | glnA |
| Synonyms | |
| Essential | no |
| Product | trigger enzyme: glutamine synthetase |
| Function | glutamine biosynthesis, control of TnrA and GlnR activity |
| Metabolic function and regulation of this protein in SubtiPathways: Ammonium/ glutamate | |
| MW, pI | 50 kDa, 4.874 |
| Gene length, protein length | 1332 bp, 444 aa |
| Immediate neighbours | glnR, ynxB |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 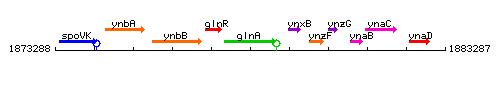
This image was kindly provided by SubtiList
| |
Contents
The gene
Basic information
- Locus tag: BSU17460
Phenotypes of a mutant
auxotrophic for glutamine
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: ATP + L-glutamate + NH3 = ADP + phosphate + L-glutamine (according to Swiss-Prot)
- Protein family: glutamine synthetase family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information: K(M) for: Glu: 27 mM, ATP: 2.4 mM, ammonium: 0.18 mM; v(max): 3.7 µmol/min/mg
- Domains: glutamate binding flap (aa 300 ... 306: protects unstable intermediates from abberant hydrolysis)
- Modification: phosphorylated on ser/ thr/ tyr PubMed, in vitro phosphorylated by PrkC on Thr-26, Thr-147, Ser-207, and Thr-286 PubMed
- Cofactor(s): Mg(2+)
- Effectors of protein activity: feedback inhibition by glutamine, glutamine binds the entrance site for glutamate
- Interactions: TnrA-GlnA PubMed, GlnR-GlnA PubMed, (only the feedback-inhibited enzyme interacts with TnrA and GlnR)
- Localization: cytoplasm (according to Swiss-Prot)
Database entries
- Structure: 2GLS (enzyme from Salmonella typhimurium)
- UniProt: P12425
- KEGG entry: [3]
- E.C. number: 6.3.1.2
Additional information
GlnA is a homooligomer of 12 subunits
Expression and regulation
- Regulation:
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant: GP247 (cat), available in Stülke lab
- Expression vector:
- GFP fusion:
- two-hybrid system:
- Antibody: available in Karl Forchhammer lab
Labs working on this gene/protein
Susan Fisher, Boston, USA homepage
Your additional remarks
References
Reviews
Fabian M Commichau, Jörg Stülke
Trigger enzymes: bifunctional proteins active in metabolism and in controlling gene expression.
Mol Microbiol: 2008, 67(4);692-702
[PubMed:18086213]
[WorldCat.org]
[DOI]
(P p)
S H Fisher
Regulation of nitrogen metabolism in Bacillus subtilis: vive la différence!
Mol Microbiol: 1999, 32(2);223-32
[PubMed:10231480]
[WorldCat.org]
[DOI]
(P p)
Original publications