Difference between revisions of "DegU"
(→Extended information on the protein) |
|||
| Line 82: | Line 82: | ||
* '''Effectors of protein activity:''' DNA-binding activity of DegU is inhibited by [[RapG]] {{PubMed|12950930}} | * '''Effectors of protein activity:''' DNA-binding activity of DegU is inhibited by [[RapG]] {{PubMed|12950930}} | ||
| − | * '''Interactions:''' [[DegU]]-[[RapG]] {{PubMed|12950930}}, [[DegS]]-[[DegU]] | + | * '''[[SubtInteract|Interactions]]:''' |
| + | ** [[DegU]]-[[RapG]] {{PubMed|12950930}}, [[DegS]]-[[DegU]] | ||
| − | * '''Localization:''' cytoplasm (according to Swiss-Prot) | + | * '''[[Localization]]:''' cytoplasm (according to Swiss-Prot) |
=== Database entries === | === Database entries === | ||
Revision as of 12:24, 10 August 2011
- Description: two-component response regulator, regulation of degradative enzyme and genetic competence, involved in activation of capsule biosynthetic operon expression (together with SwrAA), non-phosphorylated DegU is required for swarming motility PubMed
| Gene name | degU |
| Synonyms | sacU, iep |
| Essential | no |
| Product | two-component response regulator |
| Function | regulation of degradative enzyme and genetic competence |
| Interactions involving this protein in SubtInteract: DegU | |
| Metabolic function and regulation of this protein in SubtiPathways: Protein secretion | |
| MW, pI | 25 kDa, 5.602 |
| Gene length, protein length | 687 bp, 229 aa |
| Immediate neighbours | yviA, degS |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 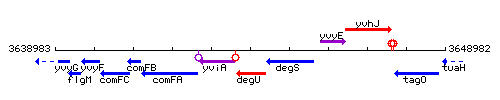
This image was kindly provided by SubtiList
| |
Contents
Categories containing this gene/protein
biofilm formation, transcription factors and their control, phosphoproteins
This gene is a member of the following regulons
The DegU regulon
The gene
Basic information
- Locus tag: BSU35490
Phenotypes of a mutant
- defect in biofilm formation PubMed, this can be suppressed by DegU-independent expression of YuaB PubMed
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: transcription activator of degradative enzyme genes (sacB and degQ) and of comK expression
- Protein family:
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Cofactor(s):
- Localization: cytoplasm (according to Swiss-Prot)
Database entries
- Structure:
- UniProt: P13800
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Reviews
Daniel López, Roberto Kolter
Extracellular signals that define distinct and coexisting cell fates in Bacillus subtilis.
FEMS Microbiol Rev: 2010, 34(2);134-49
[PubMed:20030732]
[WorldCat.org]
[DOI]
(I p)
Ewan J Murray, Taryn B Kiley, Nicola R Stanley-Wall
A pivotal role for the response regulator DegU in controlling multicellular behaviour.
Microbiology (Reading): 2009, 155(Pt 1);1-8
[PubMed:19118340]
[WorldCat.org]
[DOI]
(P p)
The DegU regulon
U Mäder, H Antelmann, T Buder, M K Dahl, M Hecker, G Homuth
Bacillus subtilis functional genomics: genome-wide analysis of the DegS-DegU regulon by transcriptomics and proteomics.
Mol Genet Genomics: 2002, 268(4);455-67
[PubMed:12471443]
[WorldCat.org]
[DOI]
(P p)
Original Publications