GapA
- Description: Glyceraldehyde 3-phosphate dehydrogenase, NAD-dependent, glycolytic enzyme, forms a transhydrogenation cycle with GapB for balancing of NADPH
| Gene name | gapA |
| Synonyms | |
| Essential | Yes (PubMed) |
| Product | glyceraldehyde 3-phosphate dehydrogenase |
| Function | catabolic enzyme in glycolysis |
| Gene expression levels in SubtiExpress: gapA | |
| Interactions involving this protein in SubtInteract: GapA | |
| Metabolic function and regulation of this protein in SubtiPathways: Central C-metabolism | |
| MW, pI | 35.7 kDa, 5.03 |
| Gene length, protein length | 1005 bp, 335 amino acids |
| Immediate neighbours | pgk, cggR |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed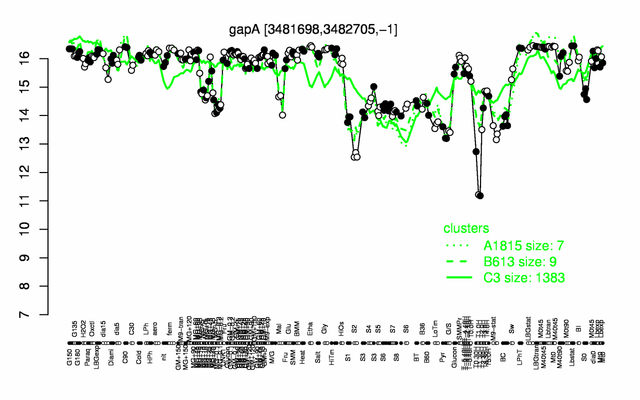
| |
Contents
Categories containing this gene/protein
carbon core metabolism, essential genes, membrane proteins, phosphoproteins
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU33940
Phenotypes of a mutant
- Essential PubMed
Database entries
- DBTBS entry: [1]
- SubtiList entry:[2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: D-glyceraldehyde 3-phosphate + phosphate + NAD+ = 3-phospho-D-glyceroyl phosphate + NADH (according to Swiss-Prot)
- This reaction is part of the glycolysis.
- Protein family: glyceraldehyde-3-phosphate dehydrogenase family (according to Swiss-Prot)
- Paralogous protein(s): GapB
Extended information on the protein
- Kinetic information: Michaelis-Menten PubMed
- Domains:
- Modification:
- Cofactor(s): NAD+ (do not accept NADP+) PubMed
- Effectors of protein activity:
Database entries
- Structure:
- UniProt: P09124
- KEGG entry: [3]
- E.C. number: 1.2.1.12
Additional information
- GAP dehydrogenases from different sources (incl. Geobacillus stearothermophilus) were shown to cleave RNA (PubMed)
- Moreover, mutations in gapA from B. subtilis can suppress mutations in genes involved in DNA replication (PubMed).
- extensive information on the structure and enzymatic properties of GapA can be found at Proteopedia
Expression and regulation
- Database entries: DBTBS
- Additional information:
- GapA is one of the most abundant proteins in the cell. In the presence of glucose, there are about 25,000 GapA molecules per cell (PubMed).
- The primary mRNAs of the operon are highly unstable. The primary mRNA is subject to processing at the very end of the cggR open reading frame. This results in stable mature gapA and gapA-pgk-tpiA-pgm-eno mRNAs. PubMed The processing event requires the RNase Y PubMed.
- The accumulation of the cggR-gapA mRNA is strongly dependent on the presence of the YkzW peptide, due to stabilization of the mRNA PubMed.
- the mRNA is substantially stabilized upon depletion of RNase Y PubMed
Biological materials
- Mutant:
- Expression vector:
- GFP fusion:
- two-hybrid system: B. pertussis adenylate cyclase-based bacterial two hybrid system (BACTH), available in Stülke lab
- Antibody: available in Stülke lab
Labs working on this gene/protein
Stephane Aymerich, Microbiology and Molecular Genetics, INRA Paris-Grignon, France
Jörg Stülke, University of Göttingen, Germany homepage
Your additional remarks
References
Additional publications: PubMed
Fabian M Commichau, Nico Pietack, Jörg Stülke
Essential genes in Bacillus subtilis: a re-evaluation after ten years.
Mol Biosyst: 2013, 9(6);1068-75
[PubMed:23420519]
[WorldCat.org]
[DOI]
(I p)
Martin Rühl, Dominique Le Coq, Stéphane Aymerich, Uwe Sauer
13C-flux analysis reveals NADPH-balancing transhydrogenation cycles in stationary phase of nitrogen-starving Bacillus subtilis.
J Biol Chem: 2012, 287(33);27959-70
[PubMed:22740702]
[WorldCat.org]
[DOI]
(I p)
Alexander K W Elsholz, Kürsad Turgay, Stephan Michalik, Bernd Hessling, Katrin Gronau, Dan Oertel, Ulrike Mäder, Jörg Bernhardt, Dörte Becher, Michael Hecker, Ulf Gerth
Global impact of protein arginine phosphorylation on the physiology of Bacillus subtilis.
Proc Natl Acad Sci U S A: 2012, 109(19);7451-6
[PubMed:22517742]
[WorldCat.org]
[DOI]
(I p)
Lehnik-Habrink M, Schaffer M, Mäder U, Diethmaier C, Herzberg C, Stülke J RNA processing in Bacillus subtilis: identification of targets of the essential RNase Y. Mol Microbiol. 2011 81(6): 1459-1473. PubMed:21815947