RemA
- Description: transcriptional regulator of the extracellular matrix genes, acts in parallel to SinR, AbrB, and DegU
| Gene name | remA |
| Synonyms | ylzA |
| Essential | no |
| Product | regulator of the extracellular matrix genes |
| Function | control of biofilm formation |
| Gene expression levels in SubtiExpress: remA | |
| Metabolic function and regulation of this protein in SubtiPathways: remA | |
| MW, pI | 9 kDa, 7.178 |
| Gene length, protein length | 267 bp, 89 aa |
| Immediate neighbours | yloC, gmk |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 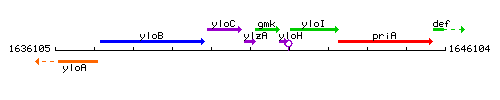
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed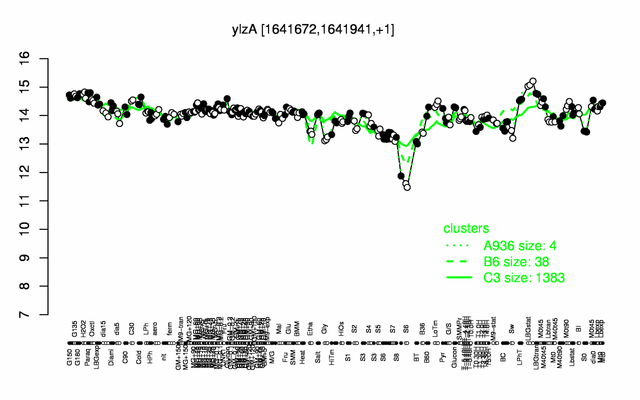
| |
Contents
Categories containing this gene/protein
transcription factors and their control, biofilm formation, sporulation proteins
This gene is a member of the following regulons
The RemA regulon
The gene
Basic information
- Locus tag: BSU15670
Phenotypes of a mutant
Database entries
- BsubCyc: BSU15670
- DBTBS entry: no entry
- SubtiList entry: [1]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family: UPF0296 family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
Database entries
- BsubCyc: BSU15670
- Structure:
- UniProt: Q7WY72
- KEGG entry: [2]
- E.C. number:
Additional information
Expression and regulation
- Operon:
- Regulation:
- Regulatory mechanism:
- Additional information:
- number of protein molecules per cell (minimal medium with glucose and ammonium): 280 PubMed
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Daniel Kearns, Indiana University, Bloomington, USA, homepage
Your additional remarks
References
Reviews
Original publications
Jared T Winkelman, Anna C Bree, Ashley R Bate, Patrick Eichenberger, Richard L Gourse, Daniel B Kearns
RemA is a DNA-binding protein that activates biofilm matrix gene expression in Bacillus subtilis.
Mol Microbiol: 2013, 88(5);984-97
[PubMed:23646920]
[WorldCat.org]
[DOI]
(I p)
Ana B Abecasis, Mónica Serrano, Renato Alves, Leonor Quintais, José B Pereira-Leal, Adriano O Henriques
A genomic signature and the identification of new sporulation genes.
J Bacteriol: 2013, 195(9);2101-15
[PubMed:23396918]
[WorldCat.org]
[DOI]
(I p)
Bjorn A Traag, Antonia Pugliese, Jonathan A Eisen, Richard Losick
Gene conservation among endospore-forming bacteria reveals additional sporulation genes in Bacillus subtilis.
J Bacteriol: 2013, 195(2);253-60
[PubMed:23123912]
[WorldCat.org]
[DOI]
(I p)
Loralyn M Cozy, Andrew M Phillips, Rebecca A Calvo, Ashley R Bate, Yi-Huang Hsueh, Richard Bonneau, Patrick Eichenberger, Daniel B Kearns
SlrA/SinR/SlrR inhibits motility gene expression upstream of a hypersensitive and hysteretic switch at the level of σ(D) in Bacillus subtilis.
Mol Microbiol: 2012, 83(6);1210-28
[PubMed:22329926]
[WorldCat.org]
[DOI]
(I p)
Jared T Winkelman, Kris M Blair, Daniel B Kearns
RemA (YlzA) and RemB (YaaB) regulate extracellular matrix operon expression and biofilm formation in Bacillus subtilis.
J Bacteriol: 2009, 191(12);3981-91
[PubMed:19363116]
[WorldCat.org]
[DOI]
(I p)